 Some of the reasons for the increased popularity of RF models are (a) they require very simple input preparation and can handle binary, categorical, count, and continuous dependent variables without the need for any preprocessing also of independent variables like scaling, (b) they perform implicit variable selection and provide a ranking of predictor (feature) importance, (c) they are inexpensive in terms of computational resources needed for their training since there are few hyperparameters that commonly need to be tuned (number of trees, number of features sampled, and number of samples in the final nodes) and because instead of working directly with all independent variables simultaneously each time, they use only a fraction of the independent variables, (d) some algorithms can beat random forests, but it is never by much, and other algorithms many times take much longer to build and tune than an RF model, (e) contrary to deep neural networks that are really hard to build, it is really hard to build a bad random forest, since it depends on very few hyperparameters and some of them are not very sensitive, which means that a lot of tweaking and fiddling is not required to get a decent random forest model, (f) they have a very simple learning algorithm, (g) they are easy to implement since there are many free and open-source implementations, and (h) RF parallelization is possible because each decision tree is grown independently.įor this reason, this chapter provides the fundamentals for building RF models as well as many illustrative examples for continuous, binary, categorical, and count response variables in the context of genomic prediction. 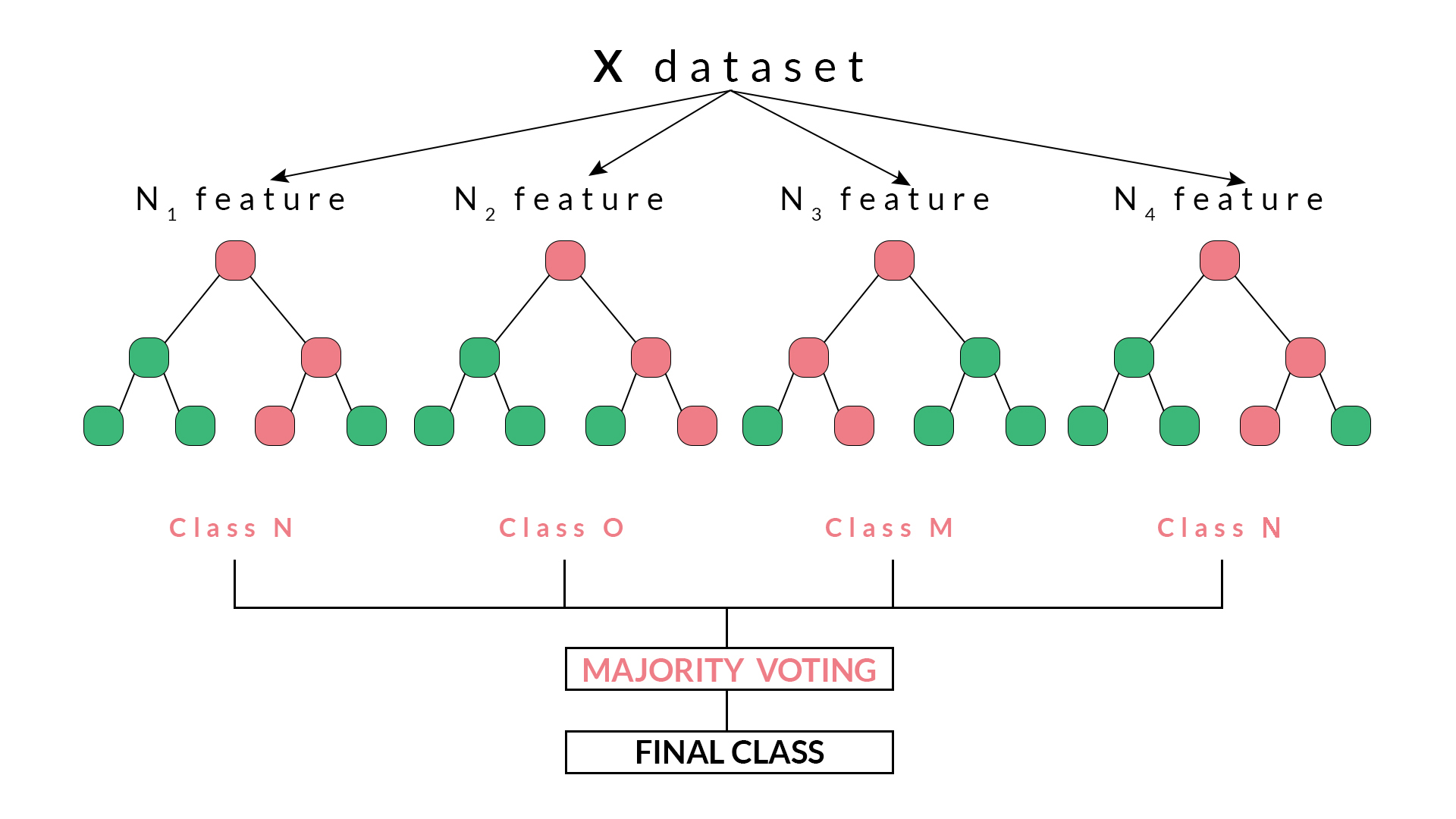
They also reported that RF produced the most consistent results with very good predictive ability and outperformed other methods in terms of correct classification. González-Recio and Forni ( 2011) found that RF performed better than Bayesian regression when detecting resistant and susceptible animals based on genetic markers. ( 2016) found that, for binary traits, RF outperformed the GBLUP method only in a scenario combining the highest heritability, the largest dense marker panel (50K SNP chip), and the largest number of QTL. ( 2009) found that RF performs better than other methods for binary traits when the sample size is large and the percentage of missing data is low (García-Magariños et al. RF is one of the models adopted for genomic prediction with many successful applications (Sarkar et al. For these reasons, RF is one of the most popular and powerful machine learning algorithms that has been successfully applied in fields such as banking, medicine, electronic commerce, stock market, and finance, among others.ĭue to the fact that there is no universal model that works in all circumstances, many statistical machine learning models have been adopted for genomic prediction. Also, RF allows measuring the relative importance of each predictor (independent variable) for the prediction. RF is a supervised machine learning algorithm that is very flexible, easy to use, and that without a lot of effort produces very competitive predictions of continuous, binary, categorical, and count outcomes. Random forest (RF) has proven to be an effective tool for such settings, already having produced numerous successful applications (Chen and Ishwaran 2012). The complexity and high dimensionality of genomic data require using flexible and powerful statistical machine learning tools for effective statistical analysis. Final comments about the pros and cons of random forest are provided. In this case, some examples are provided for illustrating its implementation even with mixed outcomes (continuous, binary, and categorical). The random forest algorithm for multivariate outcomes is provided and its most popular splitting rules are also explained. In addition, many examples are provided for training random forest models with different types of response variables with plant breeding data. 
We give (1) the random forest algorithm, (2) the main hyperparameters that need to be tuned, and (3) different splitting rules that are key for implementing random forest models for continuous, binary, categorical, and count response variables. Then we describe the process of building decision trees, which are a key component for building random forest models. The motivations for using random forest in genomic-enabled prediction are explained. We give a detailed description of random forest and exemplify its use with data from plant breeding and genomic selection. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
